Large Scale Gene Expression Data Analysis III
Yan Cui (ycui2@uthsc.edu)
The address of this webpage:
http://compbio.uthsc.edu/microarray/lecture3.htm
Algorithms
for large scale gene expression data analysis
1)
Hierarchical clustering (output: gene clusters)
2)
K-means/medians clustering (output: gene clusters)
3)
T-test (output: a group of differentially expressed genes)
4)
SAM (output: a group of differentially expressed genes)
5)
PTM (output: a group of genes matching the expression template)
What to do with these lists of selected genes?
How to understand the biological meaning of your
gene list?
Start from the existing knowledge.
First, find out what are already known about those
genes.

What is Ontology?
Ontology is the philosophical study of the nature of being, becoming, existence, or reality, as well as the basic categories of being and their relations. (from Wikipedia)
In computer science and information science, an ontology formally represents
knowledge as a set of concepts within a domain, using a shared vocabulary to denote the types, properties and
interrelationships of those concepts. (from
Wikipedia)
Gene Ontology: A system of controlled vocabulary to describe gene functions and their relations

Making
biological knowledge computable
Gene Ontology does not provide new biological
knowledge, but it is a new way of representing and organizing existing
biological knowledge. It is a standardized
and computer-readable gene function classification system.
Gene Ontology has a hierarchical structure, which is
a Directed Acyclic Graph (DAG).
Directed: A parent node and a child node are linked
by a directed edge (arrow) starting from the parent node and ending at the
child node. The parent node represents a broader functional category while the
child node represents a more specific functional category.
Acyclic: No directed cycle. A node cannot be the
descendant of itself.
Unified Gene Function Classification System for All
Organisms
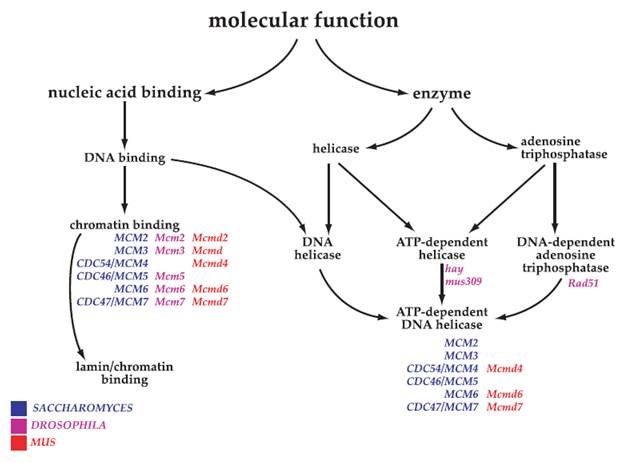
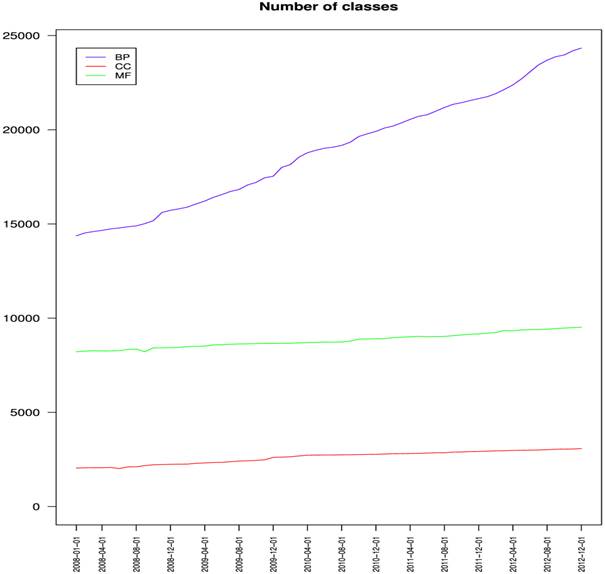
The number of classes in the three Ontologies
(from Dameron O, Bettembourg C, Le Meur N (2013) Measuring the Evolution of Ontology Complexity: The Gene Ontology Case Study. PLoS ONE 8: e75993)
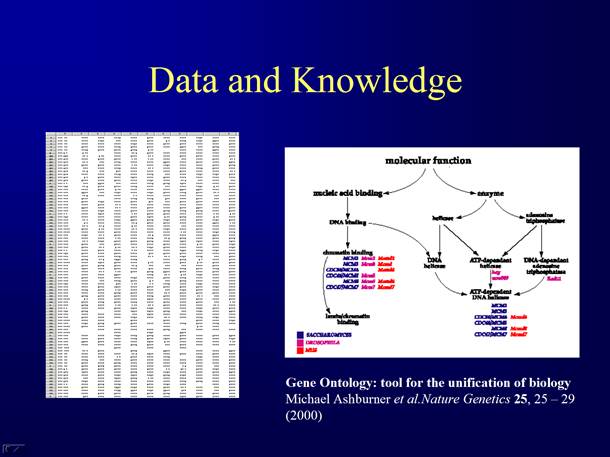
Computational tools for Gene Ontology Analysis
Help you understand the gene lists from large scale gene expression analysis.
Link results from new data to existing knowledge
Gene list
Potential markers for the prognosis of breast cancer, a set of genes differentially expressed between two groups of breast cancer patients with significantly different five year survival after surgery and chemotherapy.
NCBI Entrez Gene ID
1942 1956 2064 2625 3002 4485 4602 4605 4609 5327 6659 6662 6722 7494
Gene Symbols
EFNA1 EGFR ERBB2 GATA3 GZMB MST1 MYB MYBL2 MYC PLAT SOX4 SOX9 SRF XBP1
DAVID
Functional Annotation Tool
DAVID: Database for Annotation, Visualization, and
Integrated Discovery, a web-based tool to perform functional analysis of lists
of genes derived from genomic studies.
The wide-range collection of heterogeneous
functional annotations in the DAVID Knowledgebase
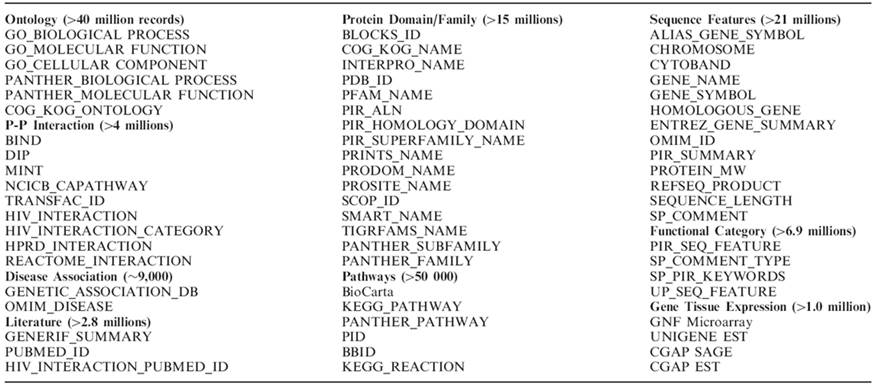

Form Huang DW et al. (2007) DAVID Bioinformatics Resources: Expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 35: W169-75.
Statistical models of enrichment (or
over-representation)
|
Selected genes |
Non-selected genes |
|
|
Having function F |
N1 |
N2 |
|
Not having function F |
N3 |
N4 |
1. Fisher Exact Probability
The Fisher exact probability for enrichment is calculated
using the Gaussian hypergeometric distribution:
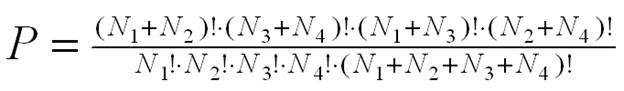 .
.
A small Fisher exact probability (e.g. <0.05)
means the association between the gene list and function F is statistically significant, therefore, function F is a biological theme of your gene list.
2. EASE Score (Adjusted Fisher Exact Probability)
The EASE score is a conservative adjustment to the
Fisher exact probability. It weights significance in favor of the association supported
by more genes.
For example, (Douglas A Hosack,
et al. Identifying biological themes within lists of genes with
EASE. Genome Biology 2003, 4:R70).
206 genes is selected from a microarray of 13,679
genes,
Only one gene in the microarray
belongs to a function category, X,
And that gene happens to appear on the
list of the 206 genes,
Fisher Exact Probability is significant (p = 0.015).
A larger function category, Y, with 787
members in the microarray,
20 members on the list of 206 genes,
Fisher exact probability is also significant (p = 0.015).
A biological theme based on the presence of a single
gene is not stable and is rarely interesting.
If the single gene happens to be a false positive,
then the significance of the biological theme is entirely false.
The EASE score is calculated by removing one gene
within category X from the list and calculating the resulting Fisher exact
probability for that category,
The EASE score: p = 1 for category X and p
= 0.027 for category Y,
Thus the EASE score eliminates the significance of
the 'unstable' category X while only slightly affecting the significance of the
more global theme Y.
The EASE score favors more robust biological themes
of the gene list.
Functional
Annotation Clustering
Similar function categories are grouped together
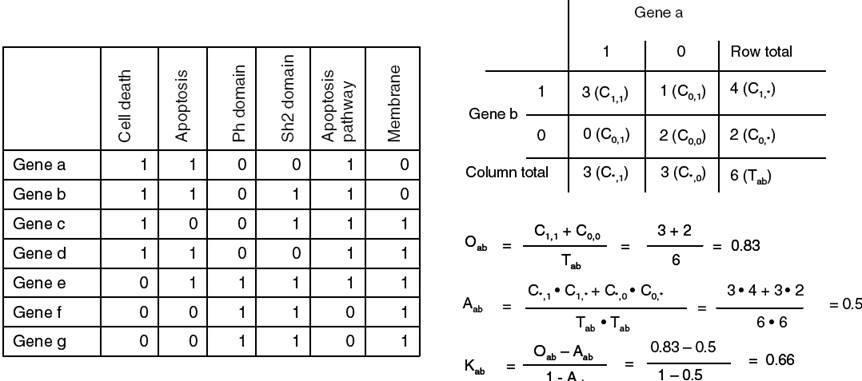
Form Huang DW et al. (2007) DAVID Bioinformatics Resources: Expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 35: W169-75.
The kappa value represents the degree of
similarity between two binary strings.
What else can you do with the DAVID Knowledgebase?
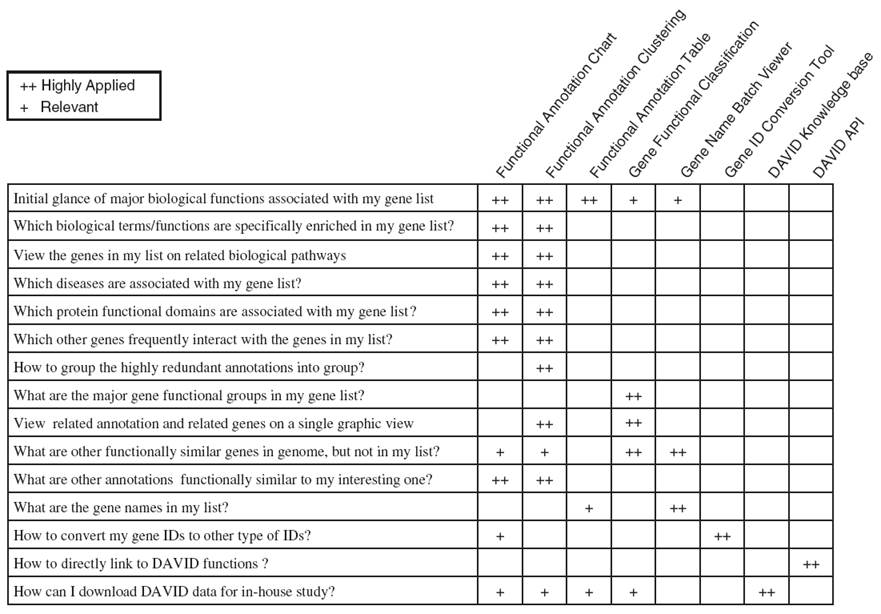
Form Huang DW et al. (2007) DAVID Bioinformatics Resources: Expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 35: W169-75.
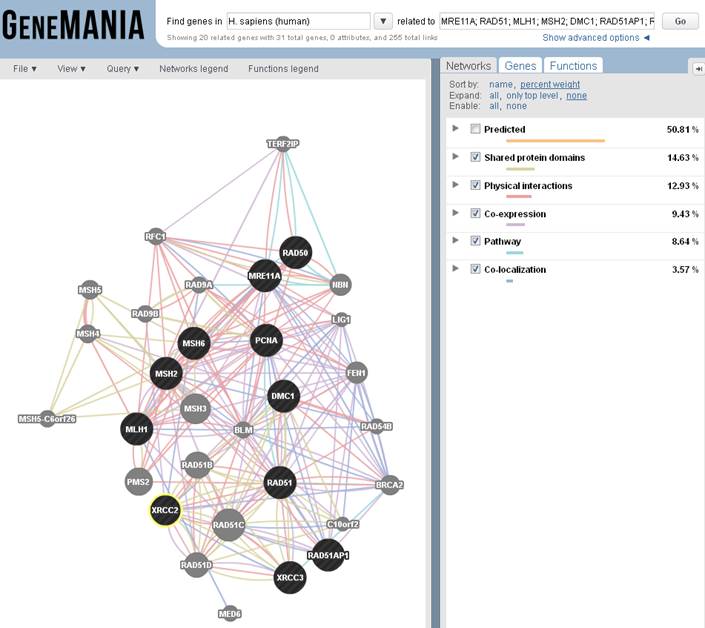
What do we know about the relationships between the
genes?
GeneMANIA uses
functional association data to connect genes to form a network.
Association data include
protein and genetic interactions, pathways, co-expression, co-localization and
protein domain similarity.
Further Reading
1.
The Gene Ontology Consortium. (2000) Gene Ontology: tool
for the unification of biology. Nat Genet 25: 25-29.
2.
The Gene Ontology Consortium. (2013) Gene Ontology
Annotations and Resources. Nucl. Acids Res. 41:
D530-D535.
3.
Huang DW, et al. (2007) DAVID Gene Functional Classification Tool: A
novel biological module-centric algorithm to functionally analyze large gene
list. Genome Biol. 8:R183.
4.
Huang, D. W., B. T. Sherman, and R. A. Lempicki.
(2008) Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat. Protocols 4: 44-57.
5.
Zuberi K, et al. (2013) GeneMANIA
Prediction Server 2013 Update. Nucl. Acids Res. 41:
W115-W122.
Homework
Due Date: Wednesday, March 5. Submit to Dr. Yan Cui via email
(ycui2@uthsc.edu).
(1) Use DAVID to identify the
major biological functions associated with the list of Affymetrix
microarray probe-sets (mouse data). What is the most significant functional
term (i.e. the term with the smallest p-value) for the gene list? How many
genes in the gene list are annotated to this term? What is the enrichment score
of the most significant annotation cluster?
102413_at
92910_at
102713_at
103445_at
104590_at
160912_i_at
102265_at
93075_r_at
103048_at
92933_at
94325_at
99041_at
101529_g_at
98981_s_at
93693_at
95460_at
(2) Use GeneMANIA to
analyze the list of potential marker genes for the prognosis of breast cancer.
Which gene is connected to GATA3 by co-localization? Which gene is connected to
PLAT by genetic interaction? What is the most significant function (i.e. the
function with the smallest FDR)?
EFNA1
EGFR
ERBB2
GATA3
GZMB
MST1
MYB
MYBL2
MYC
PLAT
SOX4
SOX9
SRF
XBP1